| |
|
Research
Ribosomes
Over the past decade, we have performed large-scale molecular dynamics
simulations of the ribosome to better understand its mechanism. Our goal is to
produce an integrated picture of tRNA selection and translocation that ties
together the large amount of existing crystallographic, cryo-EM and single
molecule FRET data. By quantifying the energy landscapes of these processes, we
are elucidating the underlying physics of protein synthesis.
Long non-coding RNAs
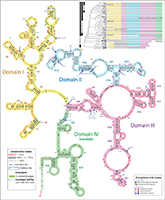
Long non-coding RNAs (lncRNAs) have emerged as an important class of RNAs in
mammalian development and disease. We are performing the first structural
studies of intact long non-coding RNAs (lncRNAs) to better understand their
mechanism. Thousands of lncRNAs have been recently identified that are
associated with stem cells, development, cancer, and brain disease. Since
several lncRNA systems have been shown to recruit epigentic factors to their
chromatin targets, lncRNAs are thought to play a key role in epigenetics.
Riboswitches
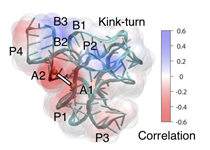
We are using a joint experiment/computation approach to understand the mechanism
of riboswitch operation. Experimentally, we are performing biochemical probing
experiments to understand the process of switching and the effect of ligand and
magnesium on this process. Computationally, we are using molecular simulation to
to understand the interplay between ligand binding, folding and magnesium
interactions on riboswitch conformation.